We’ll find a way to make these choices less confusing in a future release. It features built-in bioinformatic workflows, genome browser, sequence assembly. But of course, we use the standard definition of <0.05. Geneious Prime is a gene sequence alignment, assembly and analysis software. Some people have misunderstood this to mean that we define a single asterisk to mean P<0.0332. It shows one P value presented as “.033”, or as “0.033”, or as “0.0332” depending on the choice you made (note the difference in the number of digits and presence or absence of a leading zero). Prism 8.0-8.2 presents the choices for P value formatting like this: In this column, current versions of Prism simply write “Yes” or “No” depending on if the test corresponding to that row was found to be statistically significant or not. It would never places more than one asterisk. Prism would either places a single asterisk in that column or leaves it blank. 3.6.1 Printing 3.6.2 Saving Images 3.6.3 Exporting sequences and alignments as rich text (Prime 2019. This includes the sequence viewer, tree view, dotplot, and text view. In earlier versions of the software (Prism 6), the “Significant?” column would display a single asterisk if the t test for that row is statistically significant, given your setting for alpha and the correction for multiple comparisons. Geneious Prime allows you to print (or save as an image) the current display for any document viewer. The multiple t test analysis is different than all the rest. Note that the first two choices (APA and NEJM) show at most three asterisks (***) and the last two choices will show four asterisks with tiny P values (****). However, to use an upgraded version with more features, Geneious prime can be. P ≤ 0.0001 (For the last two choices only) Phylogenetic tree generated by Geneious a) Geneious alignment b) MUSCLE. APA (American Psychological Association) style, which shows three digits but omits the leading zero (.123). P values less than 0.001 shown as “ 0.05.Each analysis that computes P values gives you four choices: 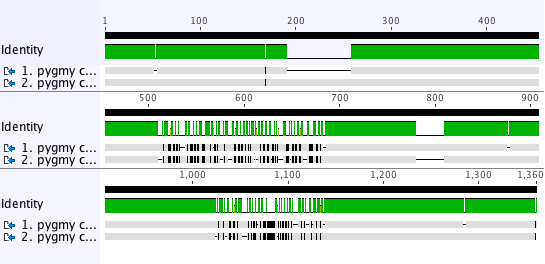
Send me an email here and I'll work on it as soon as I can.Starting with Prism 8, Prism allows you to choose which decimal format Prism will use to report P values (information on previous versions of Prism can be found below). When complete with the Pairwise Alignment tutorial, send a screen shot of the dotplot from Excercise 1.ĭetailed coverage of operations in Geneious Prime including: Information on settings, controls and options Definitions of methods and models How to run all tools in Geneious Prime and Instructions for Shared Database and Command Line Interface. That is probably the utility that Geneious. Using the associated files, learn how to align pairs of DNA and protein sequences with Geneious using dotplots and alignment algorithms. You may align the consensus sequences with multiple sequence alignment tools like Clustal Omega or MAFFT. zip Tutorial and drag (do not extract) into Geneious Prime. 
were aligned using the CLUSTALW translation alignment in Geneious Prime. Geneious Prime - Pairwise Alignment Tutorial (~30-45min)ĭownload the. Ann Arbor, MI, USA) and Geneious Prime v2019.0.4 (Biomatters Ltd., Auckland. How To Align Whole Genomes with Mauve (~5min video) Pairwise Alignment of DNA Sequences (~5min video) Pairwise Alignments - Dot Plots (~3min video) Introduction to Phylogenetic Trees (~5min video) Intro to Multiple Alignments (~2min video) Once the files have been selected, you will note that the sequences are not aligned even though all the sequences are from the same mitochondrial gene. Link to webpage with all videos: Alignment and Tree Building Learning Module 09 - Geneious Prime Alignments and PhylogeniesĪll credits to Geneious for the following videos and resources. SynBio Learn A webpage for bioinformatics resources. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
